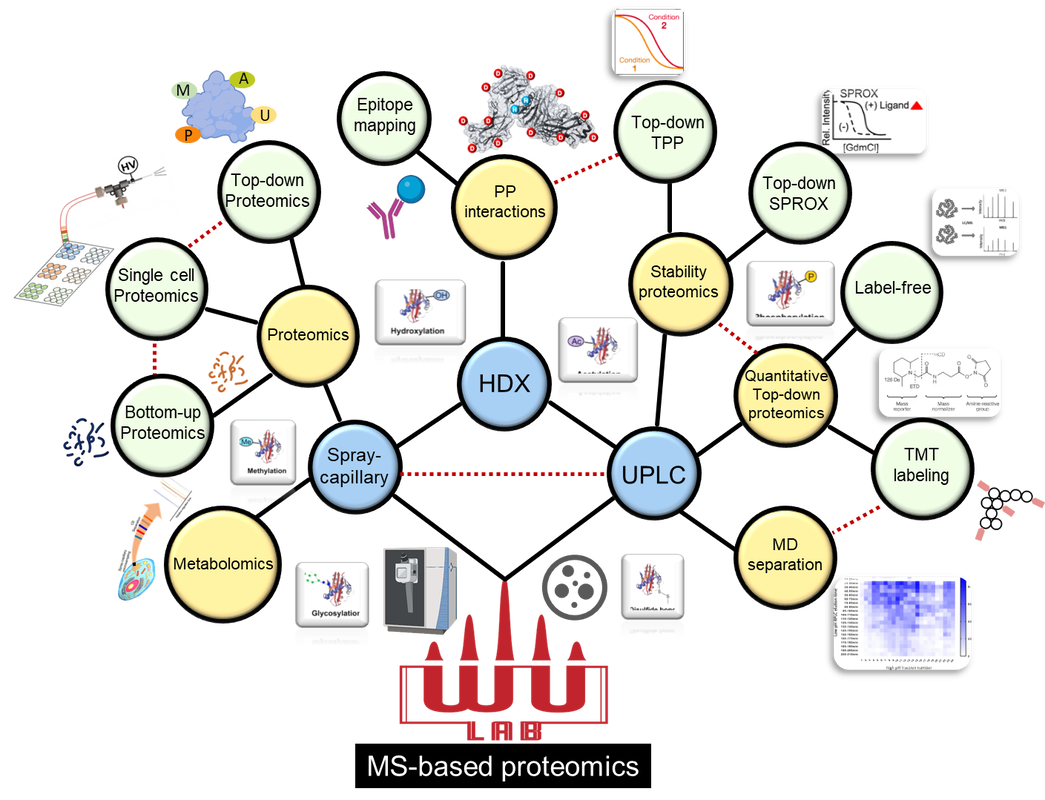
The ability to rapidly identify, characterize, and quantify proteins and understand protein functionality is critical to understanding cellular pathways and human disease. The study of human proteomics requires innovative approaches to high-throughput analysis to study these proteins in a complex and biologically relevant context. My lab focuses on the development of novel methods and technology to study and improve and high-throughput and quantitative top-down proteomics, functional proteomics, structural proteomics, and protein interactions.
High-throughput top-down proteomics: Protein PTMs have emerged in the post-genomic era as critical features in regulating and diversifying biological activity; however, identifying and characterizing proteoforms with PTMs and understanding proteoform function is currently limited by the lack of effective and accessible analytical methods. Therefore, one focus of my lab is the development of high-throughput, quantitative approaches to functionally characterize intact proteins and their modified proteoforms at the systems level. We have had success toward these goals through the development of intact protein tandem-mass-tag (TMT) labeling methods for top-down protein quantitation and high-throughput and multidimensional high-performance liquid chromatography (HPLC) techniques for deeper proteome identification. A key way we have used these technologies is in the analysis of systemic lupus erythematosus antibodies to move toward the goal of early detection and disease/symptom management.
Small-scale and single-cell omics: Cellular heterogeneity impacts cellular response to stressors and has ramifications on nearly every aspect of biomedical research. Small-scale and single-cell analysis will allow the study of phenotypically distinct cells that exist within a matrix of different cell types, data which would be indistinguishable in traditional bulk cell experiments. Single-cell omics including metabolomics and proteomics allows the quantitative analysis of metabolites and protein expression in human disease and in response to therapeutic treatment to move toward precision medicine. However, one of the most challenging aspects of single-cell analysis is small-scale sample preparation and handling. To this end, my lab has developed and patented a novel device, the ‘spray-capillary’ to perform ultralow-volume sampling and online capillary electrophoresis for sensitive and high-throughput proteomics and metabolomics. We have used the spray-capillary for omics analysis down to the single-cell level and intend to improve upon and integrate this technology in a variety of ways to study human diseases such as ovarian cancer.
Structural and functional proteomics: Research regarding protein and proteoform structure, function, and interactions is relevant to many areas of bioanalysis including the study of biological pathways, disease biomarkers and progression, antibody interactions/immunity, and therapeutic treatment/drug target discovery. High-throughput analysis of proteins in native or native-like environments is critical to analyze these complex protein systems in a meaningful way. My lab is contributing to these goals via the development of novel methods such as high-throughput hydrogen-deuterium exchange-mass spectrometry (HDX-MS) to study protein structure and probe protein-protein interactions/conformational changes and top-down stability proteomics to explore changes in proteoform functionality. Our high-throughput HDX-MS platform has been used to study antibody-antigen interactions to examine neutralizing response for human disease including anthrax and COVID-19.